
SARS-CoV-2—The Battle against Coronavirus has just Begun
•SARS-CoV-2 showed highly pathogenic, caused severe or even life-threatening diseases, and still transmitted from person-to-person.
•The World Health Organization declared a global pandemic as the coronavirus (SARS-CoV-2) rapidly spreads across the world.
•Until now, no drugs or biologics have been proven to be effective for the prevention or treatment of COVID-19.
SARS-CoV-2—Promising Antiviral Agents
Selectively and potently inhibits the RNA-dependent RNA polymerase (RdRp) of RNA viruses[1]. Shows good clinical efficacy in treating COVID-19[2].
A nucleotide analog inhibitor of RdRp. Against SARS-CoV, MERS-CoV and Ebola virus[3]. Effectively inhibits
SARS-CoV-2 in vitro[4]. Enters phase III trial.An antimalarial agent. Inhibits autophagy and toll-like receptors (TLRs)[5]. Effectively inhibits SARS-CoV-2 in vitro[6]. FDA approved.
An antimalarial and anti-inflammatory agent. Inhibits TLR7/9 signaling[7]. Efficiently inhibits SARS-CoV-2 infection in vitro[6]. FDA approved.
MedChemExpress Anti-COVID-19 Compound Library based on Relevant Proteins :
We conduct Virtual Screening of approved compound library and clinical compound library based on 3CLpro (PDB ID: 6LU7), RdRp, Spike Glycoprotein (PDB ID: 6VSB), nsp15 (PDB ID: 6VWW), PLpro and ACE2 Structure.
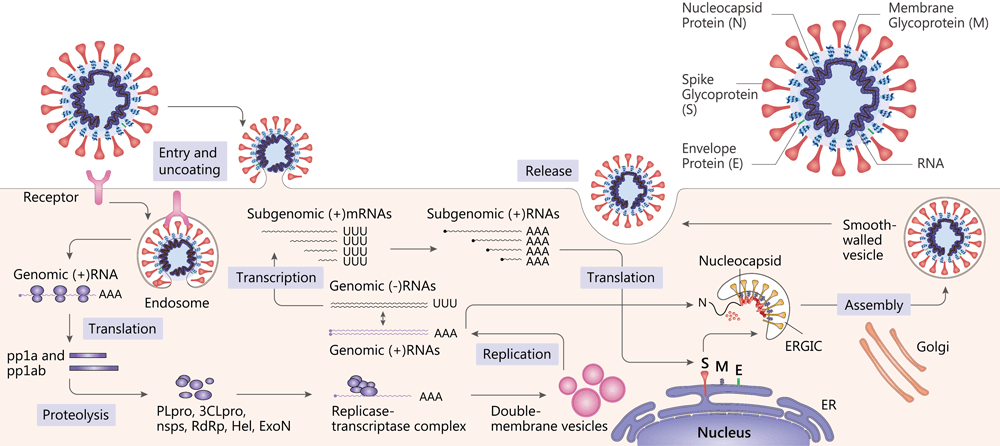
SARS-CoV and MERS-CoV structure and replication[12].
SARS-CoV-2 belongs to the Coronavirus genus in the Coronaviridae family and has a positive-sense RNA genome. Coronavirus contain four main structural proteins: spike(S), membrane (M), envelope (E), and nucleocapsid (N) proteins[8].
Attachment of the virion to the cell surface via a receptor constitutes the first step in the coronavirus life cycle[9]. SARS-CoV-2 uses the angiotensin-converting enzyme 2 (ACE2) as a cellular entry receptor[10]. Then, the virus must gain access to the host cell cytosol. This is generally accomplished by TMPRRS2 or another protease. The next step is the translation of the replicase gene from the virion genomic RNA. The replicase gene encodes two large ORFS, rep1a and rep1b, which express two co-terminal polyproteins, pp1a and pp1ab[11]. They are proteolytically cleaved into 16 non structural proteins (nsps), including papain-like protease (PLpro), 3C-like protease (3CLpro), RNA-dependent RNA polymerase (RdRp), helicase (Hel) and exonuclease (ExoN)[12].
Viral RNA synthesis follows the translation and assembly of the viral replicase complexes. It involves two stages: genome replication and subgenomic RNA transcription. Subgenomic RNAs serve as mRNAs for the structural and accessory genes which reside downstream of the replicase polyproteins[9] [11]. Following replication and subgenomic RNA synthesis, the viral structural proteins, S, E, and M are translated and inserted into the endoplasmic reticulum (ER). The mature virions are formed. Following assembly, virions are transported to the cell surface in vesicles and released by exocytosis[11].
Partial Screening Library Data :
| SARS-CoV-2 | Compounds | Information | Status |
|---|---|---|---|
| 3CLpro | Saquinavir | An HIV Protease inhibitor. | FDA approved |
| Carfilzomib | An irreversible proteasome inhibitor. | FDA approved | |
| Nelfinavir | An orally bioavailable HIV-1 protease inhibitor (Ki=2 nM) and antiviral agent. | FDA approved | |
| S Protein & ACE2 | Bimosiamose | A nonoligosaccharide pan-selectin inhibitor and has anti-inflammatory effects. | Phase 2 |
| RdRp | Zanamivir | An influenza viral neuraminidase inhibitor. | FDA approved |
| nsp15 | Ribavirin | An antiviral agent against a broad spectrum of viruses including HCV, HIV, and RSV. | FDA approved |
| PLpro | Epetraborole hydrochloride | A leucyl-tRNA synthetase (LeuRS) inhibitor. Intended for the infections caused by Gram-negative bacteria. | Phase 2 |
References:
1. Furuta Y, et al. Proc Jpn Acad Ser B Phys Biol Sci. 2017;93(7):449-463.
2. Favipiravir shows good clinical efficacy in treating COVID-19: official.
3. Agostini ML, et al. mBio. 2018 Mar 6;9(2).
4. Wang M, et al. Cell Res. 2020 Mar;30(3):269-271.
5. Mohamed FE, et al. Liver Int. 2015 Mar;35(3):1063-76.
6. Yao X, et al. Clin Infect Dis. 2020 Mar 9. pii: ciaa237.
7. Lamphier M, et al. Mol Pharmacol. 2014 Mar;85(3):429-40.
8. Guo, Y., et al. Mil Med Res. 2020 Mar 13;7(1):11.
9. Graham RL, et al. Virus Res. 2008 Apr;133(1):88-100.
10. Wanbo Tai, et al. Cell Mol Immunol. 2020 Mar 19.
11. Fehr AR, et al. Methods Mol Biol. 2015;1282:1-23.
12. de Wit E, et al. Nat Rev Microbiol. 2016 Aug;14(8):523-34.
